| |
|||
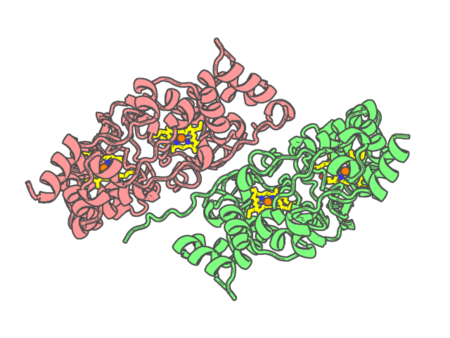 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.3 Å | |
| Deposition date | 2019-01-29 |
| |
|||
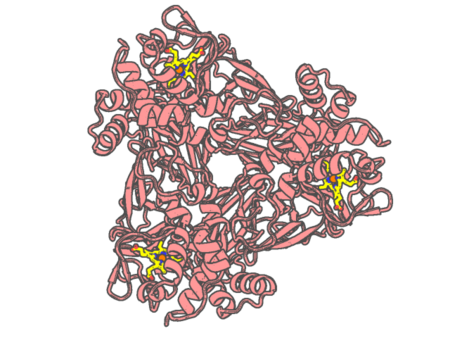 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.6 Å | |
| Deposition date | 2019-02-15 |
| |
|||
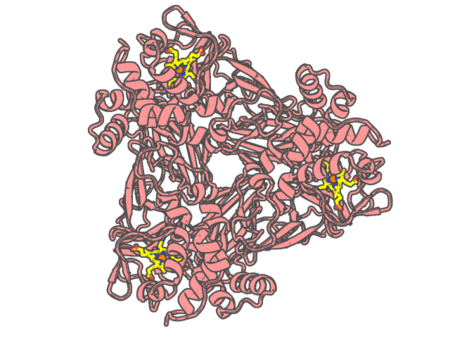 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.7 Å | |
| Deposition date | 2019-02-15 |
| |
|||
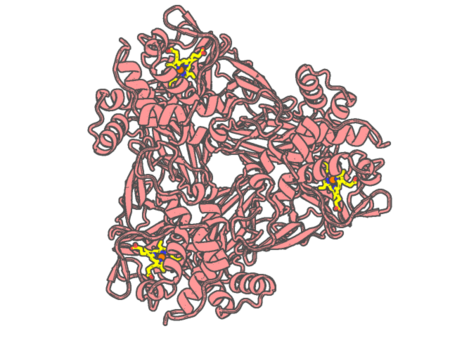 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.65 Å | |
| Deposition date | 2019-02-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.4 Å | |
| Deposition date | 2019-02-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.75 Å | |
| Deposition date | 2019-02-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.6 Å | |
| Deposition date | 2019-02-16 |
| |
|||
 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | None | |
| Deposition date | 2019-02-17 |
| |
|||
 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | None | |
| Deposition date | 2019-02-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.5 Å | |
| Deposition date | 2019-03-04 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.29 Å | |
| Deposition date | 2019-03-08 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.4 Å | |
| Deposition date | 2019-03-12 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.95 Å | |
| Deposition date | 2019-03-14 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.46 Å | |
| Deposition date | 2019-03-18 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | 2.697 Å | |
| Deposition date | 2019-03-18 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.2 Å | |
| Deposition date | 2019-03-21 |
| |
|||
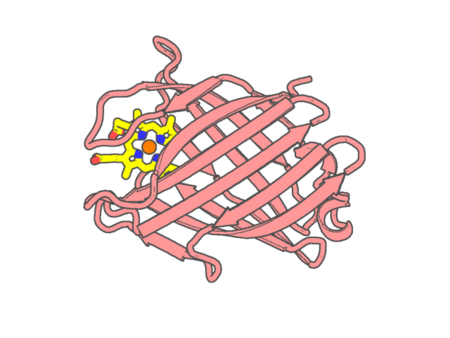 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.6 Å | |
| Deposition date | 2019-03-21 |
| |
|||
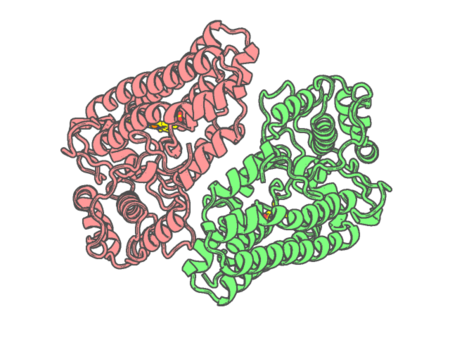 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | unavailable | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | 2.894 Å | |
| Deposition date | 2019-03-26 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.7 Å | |
| Deposition date | 2019-03-27 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2 Å | |
| Deposition date | 2019-03-27 |
