| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-03-27 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | 2.3 Å | |
| Deposition date | 2019-04-05 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | 2.64 Å | |
| Deposition date | 2019-04-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.295 Å | |
| Deposition date | 2019-04-28 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | 1.895 Å | |
| Deposition date | 2019-04-29 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 0.99 Å | |
| Deposition date | 2019-04-29 |
| |
|||
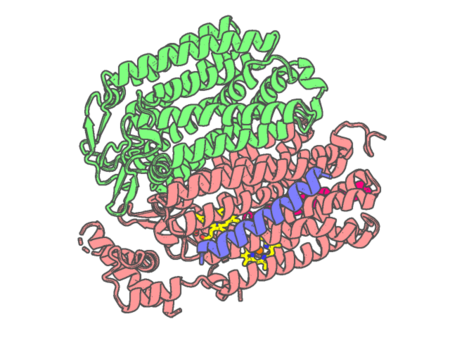 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | unavailable | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | None | |
| Deposition date | 2019-04-30 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.6 Å | |
| Deposition date | 2019-05-10 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.49 Å | |
| Deposition date | 2019-05-13 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.52 Å | |
| Deposition date | 2019-05-14 |
| |
|||
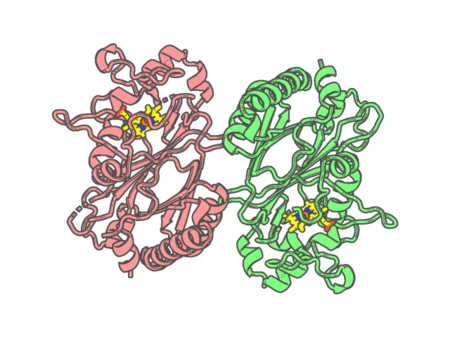 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.8 Å | |
| Deposition date | 2019-05-14 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.6 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.49 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.5 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.55 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.42 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.41 Å | |
| Deposition date | 2019-05-15 |
| |
|||
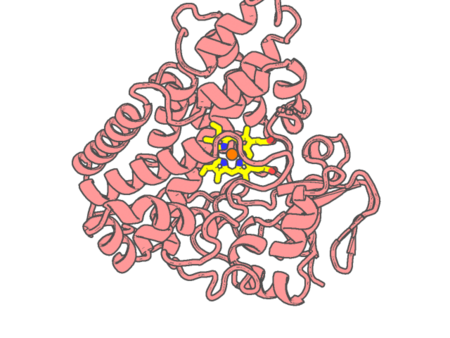 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.4 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.459 Å | |
| Deposition date | 2019-05-15 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.499 Å | |
| Deposition date | 2019-05-15 |
