| |
|||
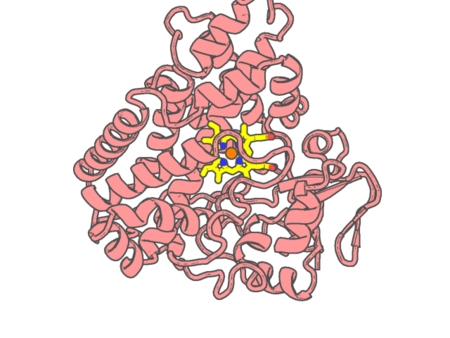 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.37 Å | |
| Deposition date | 2019-05-15 |
| |
|||
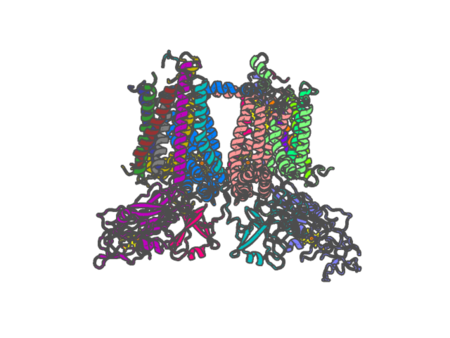 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | None | |
| Deposition date | 2019-05-15 |
| |
|||
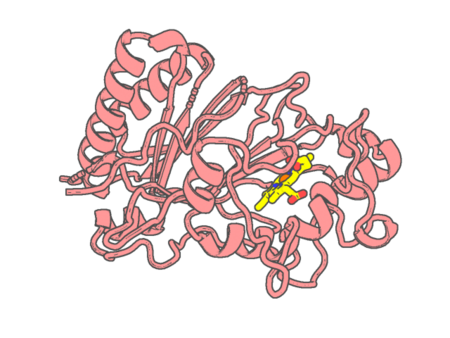 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-16 |
| |
|||
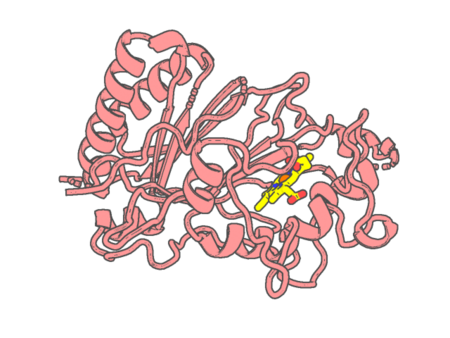 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.9 Å | |
| Deposition date | 2019-05-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.48 Å | |
| Deposition date | 2019-05-21 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.27 Å | |
| Deposition date | 2019-05-21 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.31 Å | |
| Deposition date | 2019-05-21 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.988 Å | |
| Deposition date | 2019-05-21 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.36 Å | |
| Deposition date | 2019-05-23 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.94 Å | |
| Deposition date | 2019-05-23 |
| |
|||
 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | unavailable | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | Exists / Exists | |
| Resolution | None | |
| Deposition date | 2019-06-07 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.09 Å | |
| Deposition date | 2019-07-10 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 2.502 Å | |
| Deposition date | 2019-09-16 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.75 Å | |
| Deposition date | 2019-09-17 |
| |
|||
 |
Experimental method | X-RAY DIFFRACTION |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | 1.92 Å | |
| Deposition date | 2019-09-18 |
| |
|||
 |
Experimental method | ELECTRON MICROSCOPY |
|---|---|---|
| Structural data | available | |
| Non-standard amino acids | None | |
| Missing residues/ Structural gap | None / None | |
| Resolution | None | |
| Deposition date | 2019-10-02 |
